Our Molecular Diagnostics team is now equipped with a new Next Generation Sequencing lab and we wanted to share more about it so we did a Q&A with Soledad Ulloa, our Head of Molecular Diagnostics.
Check out her answers below:
1. What is an NGS lab? What does this technology allow us?
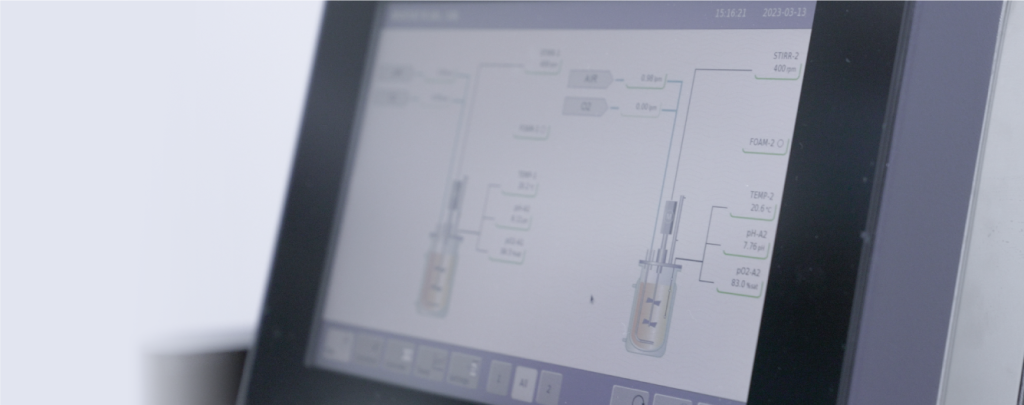
An NGS lab, or Next Generation Sequencing, is a laboratory where nucleic acid samples (DNA or RNA) are processed for later sequencing in a massive (many samples, simultaneously) and high-throughput (obtaining a large amount of data for analysis) manner.
Through NGS, we can decipher the genetic information of an organism to know the characteristics of its genome, such as genome length, virulence factors, antimicrobial resistance genes, presence of plasmids, clonality, and presence of prophages, among others. This information allows PhageLab® to develop products targeted against the infectious agent to be combated and to recommend containment and continuous improvement measures to customers.
2. How does it make our process faster and better?
At PhageLab®, the Molecular Diagnostics area has extensive equipment for molecular biology, and this new laboratory allows us to strengthen our capabilities, as we now have a dedicated space and exclusive equipment to work with the sequenced samples. It allows us to be the fastest molecular diagnostic laboratory in the agricultural industry, with a special focus on Animal Health.
3. What exactly is the Illumina technology?
Illumina technology allows deciphering genetic information by deep sequencing and high-quality short DNA fragments. The error rate of this technology is low, therefore, the sequence obtained is highly reliable. However, since these are short fragments, it is difficult to manipulate the large amount of data obtained for each sample. It is like assembling a jigsaw puzzle of small pieces without having the final image (for de novo assemblies). To finish these puzzles, our Bioinformatics team has developed specific algorithms to assemble the information and deliver a highly reliable result.
4. What does the Nanopore technology do?
Nanopore technology also allows deep sequencing, but of a lower quality, which makes the result less reliable. The advantage of this technology is the length of the sequenced fragments, which with the protocols we are using at PhageLab®, are at least 150 times longer than those sequenced by Illumina. It greatly facilitates the manipulation of the data because, following the previous analogy, if we have a puzzle made up of 2 giant pieces we do not need the reference image to assemble it.
The great advantage of having both technologies in Molecular Diagnostics is that we can now complement information and take advantage of the benefits of each one: the ease of assembling genomes using Nanopore and the high-quality sequences obtained by Illumina.
PhageLab® is registered as a trademark in Chile, Europe, Mexico, Peru and the United Kingdom, and as a word mark in Chile and Brazil.